

Version 0.7,
February 2002
Authors of
the program: Nicolas
Brouard, senior researcher at the Institut National d'Etudes
Démographiques (INED, Paris) in the
"Mortality, Health and Epidemiology" Research Unit
and Agnès
Lièvre
This program computes Healthy
Life Expectancies from cross-longitudinal data using
the methodology pioneered by Laditka and Wolf (1). Within the
family of Health Expectancies (HE), Disability-free life
expectancy (DFLE) is probably the most important index to
monitor. In low mortality countries, there is a fear that when
mortality declines, the increase in DFLE is not proportionate to
the increase in total Life expectancy. This case is called the Expansion
of morbidity. Most of the data collected today, in
particular by the international REVES
network on Health expectancy, and most HE indices based on these
data, are cross-sectional. It means that the information
collected comes from a single cross-sectional survey: people from
various ages (but mostly old people) are surveyed on their health
status at a single date. Proportion of people disabled at each
age, can then be measured at that date. This age-specific
prevalence curve is then used to distinguish, within the
stationary population (which, by definition, is the life table
estimated from the vital statistics on mortality at the same
date), the disable population from the disability-free
population. Life expectancy (LE) (or total population divided by
the yearly number of births or deaths of this stationary
population) is then decomposed into DFLE and DLE. This method of
computing HE is usually called the Sullivan method (from the name
of the author who first described it).
Age-specific proportions of people
disable are very difficult to forecast because each proportion
corresponds to historical conditions of the cohort and it is the
result of the historical flows from entering disability and
recovering in the past until today. The age-specific intensities
(or incidence rates) of entering disability or recovering a good
health, are reflecting actual conditions and therefore can be
used at each age to forecast the future of this cohort. For
example if a country is improving its technology of prosthesis,
the incidence of recovering the ability to walk will be higher at
each (old) age, but the prevalence of disability will only
slightly reflect an improve because the prevalence is mostly
affected by the history of the cohort and not by recent period
effects. To measure the period improvement we have to simulate
the future of a cohort of new-borns entering or leaving at each
age the disability state or dying according to the incidence
rates measured today on different cohorts. The proportion of
people disabled at each age in this simulated cohort will be much
lower (using the example of an improvement) that the proportions
observed at each age in a cross-sectional survey. This new
prevalence curve introduced in a life table will give a much more
actual and realistic HE level than the Sullivan method which
mostly measured the History of health conditions in this country.
Therefore, the main question is how
to measure incidence rates from cross-longitudinal surveys? This
is the goal of the IMaCH program. From your data and using IMaCH
you can estimate period HE and not only Sullivan's HE. Also the
standard errors of the HE are computed.
A cross-longitudinal survey
consists in a first survey ("cross") where individuals
from different ages are interviewed on their health status or
degree of disability. At least a second wave of interviews
("longitudinal") should measure each new individual
health status. Health expectancies are computed from the
transitions observed between waves and are computed for each
degree of severity of disability (number of life states). More
degrees you consider, more time is necessary to reach the Maximum
Likelihood of the parameters involved in the model. Considering
only two states of disability (disable and healthy) is generally
enough but the computer program works also with more health
statuses.
The simplest model is the multinomial logistic model where pij
is the probability to be observed in state j at the second
wave conditional to be observed in state i at the first
wave. Therefore a simple model is: log(pij/pii)= aij +
bij*age+ cij*sex, where 'age' is age and 'sex'
is a covariate. The advantage that this computer program claims,
comes from that if the delay between waves is not identical for
each individual, or if some individual missed an interview, the
information is not rounded or lost, but taken into account using
an interpolation or extrapolation. hPijx is the
probability to be observed in state i at age x+h
conditional to the observed state i at age x. The
delay 'h' can be split into an exact number (nh*stepm)
of unobserved intermediate states. This elementary transition (by
month or quarter trimester, semester or year) is modeled as a
multinomial logistic. The hPx matrix is simply the matrix
product of nh*stepm elementary matrices and the
contribution of each individual to the likelihood is simply hPijx.
The program presented in this
manual is a quite general program named IMaCh
(for Interpolated MArkov CHain),
designed to analyse transition data from longitudinal surveys.
The first step is the parameters estimation of a transition
probabilities model between an initial status and a final status.
From there, the computer program produces some indicators such as
observed and stationary prevalence, life expectancies and their
variances and graphs. Our transition model consists in absorbing
and non-absorbing states with the possibility of return across
the non-absorbing states. The main advantage of this package,
compared to other programs for the analysis of transition data
(For example: Proc Catmod of SAS(r)) is that the whole
individual information is used even if an interview is missing, a
status or a date is unknown or when the delay between waves is
not identical for each individual. The program can be executed
according to parameters: selection of a sub-sample, number of
absorbing and non-absorbing states, number of waves taken in
account (the user inputs the first and the last interview), a
tolerance level for the maximization function, the periodicity of
the transitions (we can compute annual, quarterly or monthly
transitions), covariates in the model. It works on Windows or on
Unix.
(1) Laditka, Sarah B. and Wolf, Douglas A. (1998), "New Methods for Analyzing Active Life Expectancy". Journal of Aging and Health. Vol 10, No. 2.
The minimum data required for a
transition model is the recording of a set of individuals
interviewed at a first date and interviewed again at least one
another time. From the observations of an individual, we obtain a
follow-up over time of the occurrence of a specific event. In
this documentation, the event is related to health status at
older ages, but the program can be applied on a lot of
longitudinal studies in different contexts. To build the data
file explained into the next section, you must have the month and
year of each interview and the corresponding health status. But
in order to get age, date of birth (month and year) is required
(missing values is allowed for month). Date of death (month and
year) is an important information also required if the individual
is dead. Shorter steps (i.e. a month) will more closely take into
account the survival time after the last interview.
In this example, 8,000 people have
been interviewed in a cross-longitudinal survey of 4 waves (1984,
1986, 1988, 1990). Some people missed 1, 2 or 3 interviews.
Health statuses are healthy (1) and disable (2). The survey is
not a real one. It is a simulation of the American Longitudinal
Survey on Aging. The disability state is defined if the
individual missed one of four ADL (Activity of daily living, like
bathing, eating, walking). Therefore, even is the individuals
interviewed in the sample are virtual, the information brought
with this sample is close to the situation of the United States.
Sex is not recorded is this sample.
Each line of the data set (named data1.txt
in this first example) is an individual record which fields are:
If your longitudinal survey do not
include information about weights or covariates, you must fill
the column with a number (e.g. 1) because a missing field is not
allowed.
This is a comment. Comments start with a '#'.
title=1st_example datafile=data1.txt lastobs=8600 firstpass=1 lastpass=4
ftol=1.e-08 stepm=1 ncov=2 nlstate=2 ndeath=1 maxwav=4 mle=1 weight=0
Intercept
and age are systematically included in the model. Additional
covariates can be included with the command
model=list of covariates
You
must write the initial guess values of the parameters for
optimisation. The number of parameters, N depends on the
number of absorbing states and non-absorbing states and on the
number of covariates.
N is given by the formula N=(nlstate +
ndeath-1)*nlstate*ncov .
Thus in the simple case with 2 covariates (the model is log
(pij/pii) = aij + bij * age where intercept and age are the two
covariates), and 2 health degrees (1 for disability-free and 2
for disability) and 1 absorbing state (3), you must enter 8
initials values, a12, b12, a13, b13, a21, b21, a23, b23. You can
start with zeros as in this example, but if you have a more
precise set (for example from an earlier run) you can enter it
and it will speed up them
Each of the four lines starts with indices "ij": ij
aij bij
# Guess values of aij and bij in log (pij/pii) = aij + bij * age
12 -14.155633 0.110794
13 -7.925360 0.032091
21 -1.890135 -0.029473
23 -6.234642 0.022315
or,
to simplify:
12 0.0 0.0
13 0.0 0.0
21 0.0 0.0
23 0.0 0.0
This
is an output if mle=1. But it can be used as
an input to get the various output data files (Health
expectancies, stationary prevalence etc.) and figures without
rerunning the rather long maximisation phase (mle=0).
The
scales are small values for the evaluation of numerical
derivatives. These derivatives are used to compute the hessian
matrix of the parameters, that is the inverse of the covariance
matrix, and the variances of health expectancies. Each line
consists in indices "ij" followed by the initial scales
(zero to simplify) associated with aij and bij.
# Scales (for hessian or gradient estimation)
12 0. 0.
13 0. 0.
21 0. 0.
23 0. 0.
This
is an output if mle=1. But it can be used as
an input to get the various output data files (Health
expectancies, stationary prevalence etc.) and figures without
rerunning the rather long maximisation phase (mle=0).
Each
line starts with indices "ijk" followed by the
covariances between aij and bij:
121 Var(a12)
122 Cov(b12,a12) Var(b12)
...
232 Cov(b23,a12) Cov(b23,b12) ... Var (b23)
# Covariance matrix
121 0.
122 0. 0.
131 0. 0. 0.
132 0. 0. 0. 0.
211 0. 0. 0. 0. 0.
212 0. 0. 0. 0. 0. 0.
231 0. 0. 0. 0. 0. 0. 0.
232 0. 0. 0. 0. 0. 0. 0. 0.
agemin=70 agemax=100 bage=50 fage=100
Once
we obtained the estimated parameters, the program is able to
calculated stationary prevalence, transitions probabilities and
life expectancies at any age. Choice of age range is useful for
extrapolation. In our data file, ages varies from age 70 to 102.
Setting bage=50 and fage=100, makes the program computing life
expectancy from age bage to age fage. As we use a model, we can
compute life expectancy on a wider age range than the age range
from the data. But the model can be rather wrong on big
intervals.
Similarly,
it is possible to get extrapolated stationary prevalence by age
ranging from agemin to agemax.
begin-prev-date=1/1/1984 end-prev-date=1/6/1988
Statements
'begin-prev-date' and 'end-prev-date' allow to select the period
in which we calculate the observed prevalences in each state. In
this example, the prevalences are calculated on data survey
collected between 1 January 1984 and 1 June 1988.
pop_based=0
The
user has the possibility to choose between population-based or
status-based health expectancies. If pop_based=0 then
status-based health expectancies are computed and if pop_based=1,
the programme computes population-based health expectancies.
Health expectancies are weighted averages of health expectancies
respective of the initial state. For a status-based index, the
weights are the cross-sectional prevalences observed between two
dates, as previously explained, whereas
for a population-based index, the weights are the stationary
prevalences.
starting-proj-date=1/1/1989 final-proj-date=1/1/1992 mov_average=0
Prevalence
and population projections are available only if the
interpolation unit is a month, i.e. stepm=1. The programme
estimates the prevalence in each state at a precise date
expressed in day/month/year. The programme computes one
forecasted prevalence a year from a starting date (1 January of
1989 in this example) to a final date (1 January 1992). The
statement mov_average allows to compute smoothed forecasted
prevalences with a five-age moving average centred at the mid-age
of the five-age period.
popforecast=0 popfile=pyram.txt popfiledate=1/1/1989 last-popfiledate=1/1/1992
This
command is available if the interpolation unit is a month, i.e.
stepm=1 and if popforecast=1. From a data file including age and
number of persons alive at the precise date ‘popfiledate’,
you can forecast the number of persons in each state until date
‘last-popfiledate’. In this example, the popfile pyram.txt includes real
data which are the Japanese population in 1989.
We
assume that you entered your 1st_example
parameter file as explained above. To
run the program you should click on the imach.exe icon and enter
the name of the parameter file which is for example C:\usr\imach\mle\biaspar.txt (you
also can click on the biaspar.txt icon located in C:\usr\imach\mle and put it with the mouse on
the imach window).
The
time to converge depends on the step unit that you used (1 month
is cpu consuming), on the number of cases, and on the number of
variables.
The
program outputs many files. Most of them are files which will be
plotted for better understanding.
Once
the optimization is finished, some graphics can be made with a
grapher. We use Gnuplot which is an interactive plotting program
copyrighted but freely distributed. A gnuplot reference manual is
available here.
When the running is finished, the user should enter a character
for plotting and output editing.
These
characters are:
The
first line is the title and displays each field of the file. The
first column is age. The fields 2 and 6 are the proportion of
individuals in states 1 and 2 respectively as observed during the
first exam. Others fields are the numbers of people in states 1,
2 or more. The number of columns increases if the number of
states is higher than 2.
The header of the file is
# Age Prev(1) N(1) N Age Prev(2) N(2) N
70 1.00000 631 631 70 0.00000 0 631
71 0.99681 625 627 71 0.00319 2 627
72 0.97125 1115 1148 72 0.02875 33 1148
It
means that at age 70, the prevalence in state 1 is 1.000 and in
state 2 is 0.00 . At age 71 the number of individuals in state 1
is 625 and in state 2 is 2, hence the total number of people aged
71 is 625+2=627.
This
file contains all the maximisation results:
-2 log likelihood= 21660.918613445392
Estimated parameters: a12 = -12.290174 b12 = 0.092161
a13 = -9.155590 b13 = 0.046627
a21 = -2.629849 b21 = -0.022030
a23 = -7.958519 b23 = 0.042614
Covariance matrix: Var(a12) = 1.47453e-001
Var(b12) = 2.18676e-005
Var(a13) = 2.09715e-001
Var(b13) = 3.28937e-005
Var(a21) = 9.19832e-001
Var(b21) = 1.29229e-004
Var(a23) = 4.48405e-001
Var(b23) = 5.85631e-005
By
substitution of these parameters in the regression model, we
obtain the elementary transition probabilities:
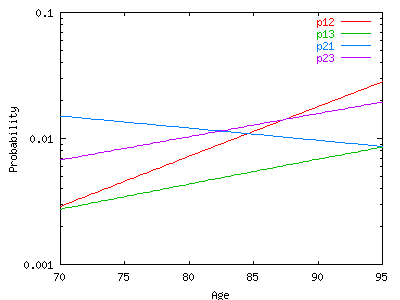
Here
are the transitions probabilities Pij(x, x+nh) where nh is a
multiple of 2 years. The first column is the starting age x (from
age 50 to 100), the second is age (x+nh) and the others are the
transition probabilities p11, p12, p13, p21, p22, p23. For
example, line 5 of the file is:
100 106 0.02655 0.17622 0.79722 0.01809 0.13678 0.84513
and
this means:
p11(100,106)=0.02655
p12(100,106)=0.17622
p13(100,106)=0.79722
p21(100,106)=0.01809
p22(100,106)=0.13678
p22(100,106)=0.84513
#Prevalence
#Age 1-1 2-2
#************
70 0.90134 0.09866
71 0.89177 0.10823
72 0.88139 0.11861
73 0.87015 0.12985
At
age 70 the stationary prevalence is 0.90134 in state 1 and
0.09866 in state 2. This stationary prevalence differs from
observed prevalence. Here is the point. The observed prevalence
at age 70 results from the incidence of disability, incidence of
recovery and mortality which occurred in the past of the cohort.
Stationary prevalence results from a simulation with actual
incidences and mortality (estimated from this cross-longitudinal
survey). It is the best predictive value of the prevalence in the
future if "nothing changes in the future". This is
exactly what demographers do with a Life table. Life expectancy
is the expected mean time to survive if observed mortality rates
(incidence of mortality) "remains constant" in the
future.
The
stationary prevalence has to be compared with the observed
prevalence by age. But both are statistical estimates and
subjected to stochastic errors due to the size of the sample, the
design of the survey, and, for the stationary prevalence to the
model used and fitted. It is possible to compute the standard
deviation of the stationary prevalence at each age.
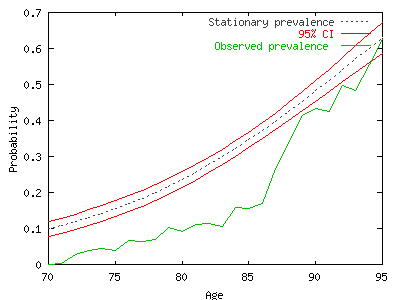
This
graph exhibits the stationary prevalence in state (2) with the
confidence interval in red. The green curve is the observed
prevalence (or proportion of individuals in state (2)). Without
discussing the results (it is not the purpose here), we observe
that the green curve is rather below the stationary prevalence.
It suggests an increase of the disability prevalence in the
future.
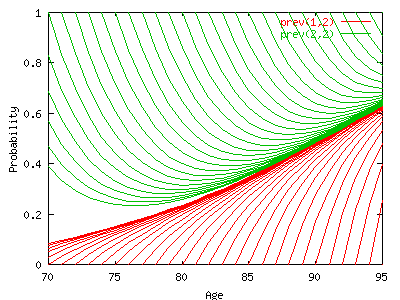
This
graph plots the conditional transition probabilities from an
initial state (1=healthy in red at the bottom, or 2=disable in
green on top) at age x to the final state 2=disable at
age x+h. Conditional means at the condition to be alive
at age x+h which is hP12x + hP22x. The
curves hP12x/(hP12x + hP22x) and hP22x/(hP12x
+ hP22x) converge with h, to the stationary
prevalence of disability. In order to get the stationary
prevalence at age 70 we should start the process at an earlier
age, i.e.50. If the disability state is defined by severe
disability criteria with only a few chance to recover, then the
incidence of recovery is low and the time to convergence is
probably longer. But we don't have experience yet.
# Health expectancies
# Age 1-1 1-2 2-1 2-2
70 10.9226 3.0401 5.6488 6.2122
71 10.4384 3.0461 5.2477 6.1599
72 9.9667 3.0502 4.8663 6.1025
73 9.5077 3.0524 4.5044 6.0401
For example 70 10.9226 3.0401 5.6488 6.2122 means:
e11=10.9226 e12=3.0401 e21=5.6488 e22=6.2122
For
example, life expectancy of a healthy individual at age 70 is
10.92 in the healthy state and 3.04 in the disability state
(=13.96 years). If he was disable at age 70, his life expectancy
will be shorter, 5.64 in the healthy state and 6.21 in the
disability state (=11.85 years). The total life expectancy is a
weighted mean of both, 13.96 and 11.85; weight is the proportion
of people disabled at age 70. In order to get a pure period index
(i.e. based only on incidences) we use the computed or
stationary prevalence at age 70 (i.e. computed from
incidences at earlier ages) instead of the observed prevalence
(for example at first exam) (see
below).
For
example, the covariances of life expectancies Cov(ei,ej) at age
50 are (line 3)
Cov(e1,e1)=0.4776 Cov(e1,e2)=0.0488=Cov(e2,e1) Cov(e2,e2)=0.0424
#Total LEs with variances: e.. (std) e.1 (std) e.2 (std)
70 13.76 (0.22) 10.40 (0.20) 3.35 (0.14)
Thus,
at age 70 the total life expectancy, e..=13.76years is the
weighted mean of e1.=13.96 and e2.=11.85 by the stationary
prevalence at age 70 which are 0.90134 in state 1 and 0.09866 in
state 2, respectively (the sum is equal to one). e.1=10.40 is the
Disability-free life expectancy at age 70 (it is again a weighted
mean of e11 and e21). e.2=3.35 is also the life expectancy at age
70 to be spent in the disability state.
This
figure represents the health expectancies and the total life
expectancy with the confident interval in dashed curve.
Standard
deviations (obtained from the information matrix of the model) of
these quantities are very useful. Cross-longitudinal surveys are
costly and do not involve huge samples, generally a few
thousands; therefore it is very important to have an idea of the
standard deviation of our estimates. It has been a big challenge
to compute the Health Expectancy standard deviations. Don't be
confuse: life expectancy is, as any expected value, the mean of a
distribution; but here we are not computing the standard
deviation of the distribution, but the standard deviation of the
estimate of the mean.
Our
health expectancies estimates vary according to the sample size
(and the standard deviations give confidence intervals of the
estimate) but also according to the model fitted. Let us explain
it in more details.
Choosing
a model means at least two kind of choices. First we have to
decide the number of disability states. Second we have to design,
within the logit model family, the model: variables, covariables,
confounding factors etc. to be included.
More
disability states we have, better is our demographical approach
of the disability process, but smaller are the number of
transitions between each state and higher is the noise in the
measurement. We do not have enough experiments of the various
models to summarize the advantages and disadvantages, but it is
important to say that even if we had huge and unbiased samples,
the total life expectancy computed from a cross-longitudinal
survey, varies with the number of states. If we define only two
states, alive or dead, we find the usual life expectancy where it
is assumed that at each age, people are at the same risk to die.
If we are differentiating the alive state into healthy and
disable, and as the mortality from the disability state is higher
than the mortality from the healthy state, we are introducing
heterogeneity in the risk of dying. The total mortality at each
age is the weighted mean of the mortality in each state by the
prevalence in each state. Therefore if the proportion of people
at each age and in each state is different from the stationary
equilibrium, there is no reason to find the same total mortality
at a particular age. Life expectancy, even if it is a very useful
tool, has a very strong hypothesis of homogeneity of the
population. Our main purpose is not to measure differential
mortality but to measure the expected time in a healthy or
disability state in order to maximise the former and minimize the
latter. But the differential in mortality complexifies the
measurement.
Incidences
of disability or recovery are not affected by the number of
states if these states are independant. But incidences estimates
are dependant on the specification of the model. More covariates
we added in the logit model better is the model, but some
covariates are not well measured, some are confounding factors
like in any statistical model. The procedure to "fit the
best model' is similar to logistic regression which itself is
similar to regression analysis. We haven't yet been so far
because we also have a severe limitation which is the speed of
the convergence. On a Pentium III, 500 MHz, even the simplest
model, estimated by month on 8,000 people may take 4 hours to
converge. Also, the program is not yet a statistical package,
which permits a simple writing of the variables and the model to
take into account in the maximisation. The actual program allows
only to add simple variables like age+sex or age+sex+ age*sex but
will never be general enough. But what is to remember, is that
incidences or probability of change from one state to another is
affected by the variables specified into the model.
Also,
the age range of the people interviewed has a link with the age
range of the life expectancy which can be estimated by
extrapolation. If your sample ranges from age 70 to 95, you can
clearly estimate a life expectancy at age 70 and trust your
confidence interval which is mostly based on your sample size,
but if you want to estimate the life expectancy at age 50, you
should rely in your model, but fitting a logistic model on a age
range of 70-95 and estimating probabilities of transition out of
this age range, say at age 50 is very dangerous. At least you
should remember that the confidence interval given by the
standard deviation of the health expectancies, are under the
strong assumption that your model is the 'true model', which is
probably not the case.
This
copy of the parameter file can be useful to re-run the program
while saving the old output files.
First,
we have estimated the observed prevalence between 1/1/1984 and
1/6/1988. The mean date of interview (weighed average of
the interviews performed between1/1/1984 and 1/6/1988) is
estimated to be 13/9/1985, as written on the top on the file.
Then we forecast the probability to be in each state.
Example,
at date 1/1/1989 :
# StartingAge FinalAge P.1 P.2 P.3
# Forecasting at date 1/1/1989
73 0.807 0.078 0.115
Since
the minimum age is 70 on the 13/9/1985, the youngest forecasted
age is 73. This means that at age a person aged 70 at 13/9/1989
has a probability to enter state1 of 0.807 at age 73 on 1/1/1989.
Similarly, the probability to be in state 2 is 0.078 and the
probability to die is 0.115. Then, on the 1/1/1989, the
prevalence of disability at age 73 is estimated to be 0.088.
# Age P.1 P.2 P.3 [Population]
# Forecasting at date 1/1/1989
75 572685.22 83798.08
74 621296.51 79767.99
73 645857.70 69320.60
# Forecasting at date 1/1/1990
76 442986.68 92721.14 120775.48
75 487781.02 91367.97 121915.51
74 512892.07 85003.47 117282.76
From the population file, we estimate the
number of people in each state. At age 73, 645857 persons are in
state 1 and 69320 are in state 2. One year latter, 512892 are
still in state 1, 85003 are in state 2 and 117282 died before
1/1/1990.
Since
you know how to run the program, it is time to test it on your
own computer. Try for example on a parameter file named imachpar.txt which is a copy of mypar.txt
included in the subdirectory of imach, mytry. Edit it to change
the name of the data file to ..\data\mydata.txt if you don't want
to copy it on the same directory. The file mydata.txt is a
smaller file of 3,000 people but still with 4 waves.
Click
on the imach.exe icon to open a window. Answer to the question: 'Enter
the parameter file name:'
| IMACH,
Version 0.7 Enter
the parameter file name: ..\mytry\imachpar.txt |
Most
of the data files or image files generated, will use the
'imachpar' string into their name. The running time is about 2-3
minutes on a Pentium III. If the execution worked correctly, the
outputs files are created in the current directory, and should be
the same as the mypar files initially included in the directory mytry.
· Output on the screen The output screen looks like this Log file
#title=MLE datafile=..\data\mydata.txt lastobs=3000 firstpass=1 lastpass=3
ftol=1.000000e-008 stepm=24 ncov=2 nlstate=2 ndeath=1 maxwav=4 mle=1 weight=0
Total number of individuals= 2965, Agemin = 70.00, Agemax= 100.92
Warning, no any valid information for:126 line=126
Warning, no any valid information for:2307 line=2307
Delay (in months) between two waves Min=21 Max=51 Mean=24.495826
These lines give some warnings on the data file and also some raw statistics on frequencies of transitions.
Age 70 1.=230 loss[1]=3.5% 2.=16 loss[2]=12.5% 1.=222 prev[1]=94.1% 2.=14
prev[2]=5.9% 1-1=8 11=200 12=7 13=15 2-1=2 21=6 22=7 23=1
Age 102 1.=0 loss[1]=NaNQ% 2.=0 loss[2]=NaNQ% 1.=0 prev[1]=NaNQ% 2.=0
· Calculation of the hessian matrix. Wait...
12345678.12.13.14.15.16.17.18.23.24.25.26.27.28.34.35.36.37.38.45.46.47.48.56.57.58.67.68.78
Inverting the hessian to get the covariance matrix. Wait...
#Hessian matrix#
3.344e+002 2.708e+004 -4.586e+001 -3.806e+003 -1.577e+000 -1.313e+002 3.914e-001 3.166e+001
2.708e+004 2.204e+006 -3.805e+003 -3.174e+005 -1.303e+002 -1.091e+004 2.967e+001 2.399e+003
-4.586e+001 -3.805e+003 4.044e+002 3.197e+004 2.431e-002 1.995e+000 1.783e-001 1.486e+001
-3.806e+003 -3.174e+005 3.197e+004 2.541e+006 2.436e+000 2.051e+002 1.483e+001 1.244e+003
-1.577e+000 -1.303e+002 2.431e-002 2.436e+000 1.093e+002 8.979e+003 -3.402e+001 -2.843e+003
-1.313e+002 -1.091e+004 1.995e+000 2.051e+002 8.979e+003 7.420e+005 -2.842e+003 -2.388e+005
3.914e-001 2.967e+001 1.783e-001 1.483e+001 -3.402e+001 -2.842e+003 1.494e+002 1.251e+004
3.166e+001 2.399e+003 1.486e+001 1.244e+003 -2.843e+003 -2.388e+005 1.251e+004 1.053e+006
# Scales
12 1.00000e-004 1.00000e-006
13 1.00000e-004 1.00000e-006
21 1.00000e-003 1.00000e-005
23 1.00000e-004 1.00000e-005
# Covariance
1 5.90661e-001
2 -7.26732e-003 8.98810e-005
3 8.80177e-002 -1.12706e-003 5.15824e-001
4 -1.13082e-003 1.45267e-005 -6.50070e-003 8.23270e-005
5 9.31265e-003 -1.16106e-004 6.00210e-004 -8.04151e-006 1.75753e+000
6 -1.15664e-004 1.44850e-006 -7.79995e-006 1.04770e-007 -2.12929e-002 2.59422e-004
7 1.35103e-003 -1.75392e-005 -6.38237e-004 7.85424e-006 4.02601e-001 -4.86776e-003 1.32682e+000
8 -1.82421e-005 2.35811e-007 7.75503e-006 -9.58687e-008 -4.86589e-003 5.91641e-005 -1.57767e-002 1.88622e-004
# agemin agemax for lifexpectancy, bage fage (if mle==0 ie no data nor Max likelihood).
agemin=70 agemax=100 bage=50 fage=100
Computing prevalence limit: result on file 'plrmypar.txt'
Computing pij: result on file 'pijrmypar.txt'
Computing Health Expectancies: result on file 'ermypar.txt'
Computing Variance-covariance of DFLEs: file 'vrmypar.txt'
Computing Total LEs with variances: file 'trmypar.txt'
Computing Variance-covariance of Prevalence limit: file 'vplrmypar.txt'
End of Imach
Once
the running is finished, the program requires a caracter:
| Type
e to edit output files, c to start again, and q for
exiting: |
First
you should enter e to edit the master file
mypar.htm.
This
software have been partly granted by Euro-REVES, a concerted
action from the European Union. It will be copyrighted
identically to a GNU software product, i.e. program and software
can be distributed freely for non commercial use. Sources are not
widely distributed today. You can get them by asking us with a
simple justification (name, email, institute) mailto:brouard@ined.fr and mailto:lievre@ined.fr .
Latest
version (0.7 of February 2002) can be accessed at http://euroreves.ined.fr/imach